Epigenetics of Notch1 regulation in pulmonary microvascular rarefaction following extrauterine growth restriction.
Tang, LL; Zhang, LY; Lao, LJ; Hu, QY; Gu, WZ; Fu, LC; Du, LZ
Respiratory research
16
66
2015
Show Abstract
Extrauterine growth restriction (EUGR) plays an important role in the developmental origin of adult cardiovascular diseases. In an EUGR rat model, we reported an elevated pulmonary arterial pressure in adults and genome-wide epigenetic modifications in pulmonary vascular endothelial cells (PVECs). However, the underlying mechanism of the early nutritional insult that results in pulmonary vascular consequences later in life remains unclear.A rat model was used to investigate the physiological and structural effect of EUGR on early pulmonary vasculature by evaluating right ventricular systolic pressure and pulmonary vascular density in male rats. Epigenetic modifications of the Notch1 gene in PVECs were evaluated.EUGR decreased pulmonary vascular density with no significant impact on right ventricular systolic pressure at 3 weeks. Decreased transcription of Notch1 was observed both at 3 and 9 weeks, in association with decreased downstream target gene, Hes-1. Chromatin immunoprecipitation and bisulfite sequencing were performed to analyze the epigenetic modifications of the Notch1 gene promoter in PVECs. EUGR caused a significantly increased H3K27me3 in the proximal Notch1 gene promoter, and increased methylation of single CpG sites in the distal Notch1 gene promoter, both at 3 and 9 weeks.We conclude that EUGR results in decreased pulmonary vascular growth in association with decreased Notch1 in PVECs. This may be mediated by increased CpG methylation and H3K27me3 in the Notch1 gene promoter region. | | 26040933
 |
Isolation and characterization of an osmotic stress and ABA induced histone deacetylase in Arachis hygogaea.
Su, LC; Deng, B; Liu, S; Li, LM; Hu, B; Zhong, YT; Li, L
Frontiers in plant science
6
512
2015
Show Abstract
Histone acetylation, which together with histone methylation regulates gene activity in response to stress, is an important epigenetic modification. There is an increasing research focus on histone acetylation in crops, but there is no information to date in peanut (Arachis hypogaea). We showed that osmotic stress and ABA affect the acetylation of histone H3 loci in peanut seedlings by immunoblotting experiments. Using RNA-seq data for peanut, we found a RPD3/HDA1-like superfamily histone deacetylase (HDAC), termed AhHDA1, whose gene is up-regulated by PEG-induced water limitation and ABA signaling. We isolated and characterized AhHDA1 from A. hypogaea, showing that AhHDA1 is very similar to an Arabidopsis HDAC (AtHDA6) and, in recombinant form, possesses HDAC activity. To understand whether and how osmotic stress and ABA mediate the peanut stress response by epigenetics, the expression of AhHDA1 and stress-responsive genes following treatment with PEG, ABA, and the specific HDAC inhibitor trichostatin A (TSA) were analyzed. AhHDA1 transcript levels were enhanced by all three treatments, as was expression of peanut transcription factor genes, indicating that AhHDA1 might be involved in the epigenetic regulation of stress resistance genes that comprise the responses to osmotic stress and ABA. | | 26217363
 |
A comprehensive epigenome map of Plasmodium falciparum reveals unique mechanisms of transcriptional regulation and identifies H3K36me2 as a global mark of gene suppression.
Karmodiya, K; Pradhan, SJ; Joshi, B; Jangid, R; Reddy, PC; Galande, S
Epigenetics & chromatin
8
32
2015
Show Abstract
Role of epigenetic mechanisms towards regulation of the complex life cycle/pathogenesis of Plasmodium falciparum, the causative agent of malaria, has been poorly understood. To elucidate stage-specific epigenetic regulation, we performed genome-wide mapping of multiple histone modifications of P. falciparum. Further to understand the differences in transcription regulation in P. falciparum and its host, human, we compared their histone modification profiles.Our comprehensive comparative analysis suggests distinct mode of transcriptional regulation in malaria parasite by virtue of poised genes and differential histone modifications. Furthermore, analysis of histone modification profiles predicted 562 genes producing anti-sense RNAs and 335 genes having bidirectional promoter activity, which raises the intriguing possibility of RNA-mediated regulation of transcription in P. falciparum. Interestingly, we found that H3K36me2 acts as a global repressive mark and gene regulation is fine tuned by the ratio of activation marks to H3K36me2 in P. falciparum. This novel mechanism of gene regulation is supported by the fact that knockout of SET genes (responsible for H3K36 methylation) leads to up-regulation of genes with highest occupancy of H3K36me2 in wild-type P. falciparum. Moreover, virulence (var) genes are mostly poised and marked by a unique set of activation (H4ac) and repression (H3K9me3) marks, which are mutually exclusive to other Plasmodium housekeeping genes.Our study reveals unique plasticity in the epigenetic regulation in P. falciparum which can influence parasite virulence and pathogenicity. The observed differences in the histone code and transcriptional regulation in P. falciparum and its host will open new avenues for epigenetic drug development against malaria parasite. | | 26388940
 |
Nucleosome competition reveals processive acetylation by the SAGA HAT module.
Ringel, AE; Cieniewicz, AM; Taverna, SD; Wolberger, C
Proceedings of the National Academy of Sciences of the United States of America
112
E5461-70
2015
Show Abstract
The Spt-Ada-Gcn5 acetyltransferase (SAGA) coactivator complex hyperacetylates histone tails in vivo in a manner that depends upon histone 3 lysine 4 trimethylation (H3K4me3), a histone mark enriched at promoters of actively transcribed genes. SAGA contains a separable subcomplex known as the histone acetyltransferase (HAT) module that contains the HAT, Gcn5, bound to Sgf29, Ada2, and Ada3. Sgf29 contains a tandem Tudor domain that recognizes H3K4me3-containing peptides and is required for histone hyperacetylation in vivo. However, the mechanism by which H3K4me3 recognition leads to lysine hyperacetylation is unknown, as in vitro studies show no effect of the H3K4me3 modification on histone peptide acetylation by Gcn5. To determine how H3K4me3 binding by Sgf29 leads to histone hyperacetylation by Gcn5, we used differential fluorescent labeling of histones to monitor acetylation of individual subpopulations of methylated and unmodified nucleosomes in a mixture. We find that the SAGA HAT module preferentially acetylates H3K4me3 nucleosomes in a mixture containing excess unmodified nucleosomes and that this effect requires the Tudor domain of Sgf29. The H3K4me3 mark promotes processive, multisite acetylation of histone H3 by Gcn5 that can account for the different acetylation patterns established by SAGA at promoters versus coding regions. Our results establish a model for Sgf29 function at gene promoters and define a mechanism governing crosstalk between histone modifications. | | 26401015
 |
The complex pattern of epigenomic variation between natural yeast strains at single-nucleosome resolution.
Filleton, F; Chuffart, F; Nagarajan, M; Bottin-Duplus, H; Yvert, G
Epigenetics & chromatin
8
26
2015
Show Abstract
Epigenomic studies on humans and model species have revealed substantial inter-individual variation in histone modification profiles. However, the pattern of this variation has not been precisely characterized, particularly regarding which genomic features are enriched for variability and whether distinct histone marks co-vary synergistically. Yeast allows us to investigate intra-species variation at high resolution while avoiding other sources of variation, such as cell type or subtype.We profiled histone marks H3K4me3, H3K9ac, H3K14ac, H4K12ac and H3K4me1 in three unrelated wild strains of Saccharomyces cerevisiae at single-nucleosome resolution and analyzed inter-strain differences statistically. All five marks varied significantly at specific loci, but to different extents. The number of nucleosomes varying for a given mark between two strains ranged from 20 to several thousands; +1 nucleosomes were significantly less subject to variation. Genes with highly evolvable or responsive expression showed higher variability; however, the variation pattern could not be explained by known transcriptional differences between the strains. Synergistic variation of distinct marks was not systematic, with surprising differences between functionally related H3K9ac and H3K14ac. Interestingly, H3K14ac differences that persisted through transient hyperacetylation were supported by H3K4me3 differences, suggesting stabilization via cross talk.Quantitative variation of histone marks among S. cerevisiae strains is abundant and complex. Its relation to functional characteristics is modular and seems modest, with partial association with gene expression divergences, differences between functionally related marks and partial co-variation between marks that may confer stability. Thus, the specific context of studies, such as which precise marks, individuals and genomic loci are investigated, is primordial in population epigenomics studies. The complexity found in this pilot survey in yeast suggests that high complexity can be anticipated among higher eukaryotes, including humans. | | 26229551
 |
Panspecies small-molecule disruptors of heterochromatin-mediated transcriptional gene silencing.
Castonguay, E; White, SA; Kagansky, A; St-Cyr, DJ; Castillo, AG; Brugger, C; White, R; Bonilla, C; Spitzer, M; Earnshaw, WC; Schalch, T; Ekwall, K; Tyers, M; Allshire, RC
Molecular and cellular biology
35
662-74
2015
Show Abstract
Heterochromatin underpins gene repression, genome integrity, and chromosome segregation. In the fission yeast Schizosaccharomyces pombe, conserved protein complexes effect heterochromatin formation via RNA interference-mediated recruitment of a histone H3 lysine 9 methyltransferase to cognate chromatin regions. To identify small molecules that inhibit heterochromatin formation, we performed an in vivo screen for loss of silencing of a dominant selectable kanMX reporter gene embedded within fission yeast centromeric heterochromatin. Two structurally unrelated compounds, HMS-I1 and HMS-I2, alleviated kanMX silencing and decreased repressive H3K9 methylation levels at the transgene. The decrease in methylation caused by HMS-I1 and HMS-I2 was observed at all loci regulated by histone methylation, including centromeric repeats, telomeric regions, and the mating-type locus, consistent with inhibition of the histone deacetylases (HDACs) Clr3 and/or Sir2. Chemical-genetic epistasis and expression profiles revealed that both compounds affect the activity of the Clr3-containing Snf2/HDAC repressor complex (SHREC). In vitro HDAC assays revealed that HMS-I1 and HMS-I2 inhibit Clr3 HDAC activity. HMS-I1 also alleviated transgene reporter silencing by heterochromatin in Arabidopsis and a mouse cell line, suggesting a conserved mechanism of action. HMS-I1 and HMS-I2 bear no resemblance to known inhibitors of chromatin-based activities and thus represent novel chemical probes for heterochromatin formation and function. | | 25487573
 |
Insulin-response epigenetic activation of Egr-1 and JunB genes at the nuclear periphery by A-type lamin-associated pY19-Caveolin-2 in the inner nuclear membrane.
Jeong, K; Kwon, H; Lee, J; Jang, D; Pak, Y
Nucleic Acids Res
43
3114-27
2015
Show Abstract
Insulin controls transcription to sustain its physiologic effects for the organism to adapt to environmental changes added to genetic predisposition. Nevertheless, insulin-induced transcriptional regulation by epigenetic factors and in defined nuclear territory remains elusive. Here we show that inner nuclear membrane (INM)-integrated caveolin-2 (Cav-2) regulates insulin-response epigenetic activation of Egr-1 and JunB genes at the nuclear periphery. INM-targeted pY19-Cav-2 in response to insulin associates specifically with the A-type lamin, disengages the repressed Egr-1 and JunB promoters from lamin A/C through disassembly of H3K9me3, and facilitates assembly of H3K9ac, H3K18ac and H3K27ac by recruitment of GCN5 and p300 and the subsequent enrichment of RNA polymerase II (Pol II) on the promoters at the nuclear periphery. Our findings show that Cav-2 is an epigenetic regulator of histone H3 modifications, and provide novel mechanisms of insulin-response epigenetic activation at the nuclear periphery. | | 25753664
 |
3D-DIP-Chip: a microarray-based method to measure genomic DNA damage.
Powell, JR; Bennett, MR; Evans, KE; Yu, S; Webster, RM; Waters, R; Skinner, N; Reed, SH
Scientific reports
5
7975
2015
Show Abstract
Genotoxins cause DNA damage, which can result in genomic instability. The genetic changes induced have far-reaching consequences, often leading to diseases such as cancer. A wide range of genotoxins exists, including radiations and chemicals found naturally in the environment, and in man-made forms created by human activity across a variety of industries. Genomic technologies offer the possibility of unravelling the mechanisms of genotoxicity, including the repair of genetic damage, enhancing our ability to develop, test and safely use existing and novel materials. We have developed 3D-DIP-Chip, a microarray-based method to measure the prevalence of genomic genotoxin-induced DNA damage. We demonstrate the measurement of both physical and chemical induced DNA damage spectra, integrating the analysis of these with the associated changes in histone acetylation induced in the epigenome. We discuss the application of the method in the context of basic and translational sciences. | | 25609656
 |
A PWWP domain-containing protein targets the NuA3 acetyltransferase complex via histone H3 lysine 36 trimethylation to coordinate transcriptional elongation at coding regions.
Gilbert, TM; McDaniel, SL; Byrum, SD; Cades, JA; Dancy, BC; Wade, H; Tackett, AJ; Strahl, BD; Taverna, SD
Molecular & cellular proteomics : MCP
13
2883-95
2014
Show Abstract
Post-translational modifications of histones, such as acetylation and methylation, are differentially positioned in chromatin with respect to gene organization. For example, although histone H3 is often trimethylated on lysine 4 (H3K4me3) and acetylated on lysine 14 (H3K14ac) at active promoter regions, histone H3 lysine 36 trimethylation (H3K36me3) occurs throughout the open reading frames of transcriptionally active genes. The conserved yeast histone acetyltransferase complex, NuA3, specifically binds H3K4me3 through a plant homeodomain (PHD) finger in the Yng1 subunit, and subsequently catalyzes the acetylation of H3K14 through the histone acetyltransferase domain of Sas3, leading to transcription initiation at a subset of genes. We previously found that Ylr455w (Pdp3), an uncharacterized proline-tryptophan-tryptophan-proline (PWWP) domain-containing protein, copurifies with stable members of NuA3. Here, we employ mass-spectrometric analysis of affinity purified Pdp3, biophysical binding assays, and genetic analyses to classify NuA3 into two functionally distinct forms: NuA3a and NuA3b. Although NuA3a uses the PHD finger of Yng1 to interact with H3K4me3 at the 5'-end of open reading frames, NuA3b contains the unique member, Pdp3, which regulates an interaction between NuA3b and H3K36me3 at the transcribed regions of genes through its PWWP domain. We find that deletion of PDP3 decreases NuA3-directed transcription and results in growth defects when combined with transcription elongation mutants, suggesting NuA3b acts as a positive elongation factor. Finally, we determine that NuA3a, but not NuA3b, is synthetically lethal in combination with a deletion of the histone acetyltransferase GCN5, indicating NuA3b has a specialized role at coding regions that is independent of Gcn5 activity. Collectively, these studies define a new form of the NuA3 complex that associates with H3K36me3 to effect transcriptional elongation. MS data are available via ProteomeXchange with identifier PXD001156. | | 25104842
 |
Comprehensive analysis of interacting proteins and genome-wide location studies of the Sas3-dependent NuA3 histone acetyltransferase complex.
Vicente-Muñoz, S; Romero, P; Magraner-Pardo, L; Martinez-Jimenez, CP; Tordera, V; Pamblanco, M
FEBS open bio
4
996-1006
2014
Show Abstract
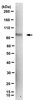
Histone acetylation affects several aspects of gene regulation, from chromatin remodelling to gene expression, by modulating the interplay between chromatin and key transcriptional regulators. The exact molecular mechanism underlying acetylation patterns and crosstalk with other epigenetic modifications requires further investigation. In budding yeast, these epigenetic markers are produced partly by histone acetyltransferase enzymes, which act as multi-protein complexes. The Sas3-dependent NuA3 complex has received less attention than other histone acetyltransferases (HAT), such as Gcn5-dependent complexes. Here, we report our analysis of Sas3p-interacting proteins using tandem affinity purification (TAP), coupled with mass spectrometry. This analysis revealed Pdp3p, a recently described component of NuA3, to be one of the most abundant Sas3p-interacting proteins. The PDP3 gene, was TAP-tagged and protein complex purification confirmed that Pdp3p co-purified with the NuA3 protein complex, histones, and several transcription-related and chromatin remodelling proteins. Our results also revealed that the protein complexes associated with Sas3p presented HAT activity even in the absence of Gcn5p and vice versa. We also provide evidence that Sas3p cannot substitute Gcn5p in acetylation of lysine 9 in histone H3 in vivo. Genome-wide occupancy of Sas3p using ChIP-on-chip tiled microarrays showed that Sas3p was located preferentially within the 5'-half of the coding regions of target genes, indicating its probable involvement in the transcriptional elongation process. Hence, this work further characterises the function and regulation of the NuA3 complex by identifying novel post-translational modifications in Pdp3p, additional Pdp3p-co-purifying chromatin regulatory proteins involved in chromatin-modifying complex dynamics and gene regulation, and a subset of genes whose transcriptional elongation is controlled by this complex. | Western Blotting | 25473596
 |