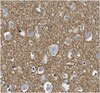
Transcription of the transforming growth factor beta activating integrin beta8 subunit is regulated by SP3, AP-1, and the p38 pathway.
Markovics, JA; Araya, J; Cambier, S; Jablons, D; Hill, A; Wolters, PJ; Nishimura, SL
The Journal of biological chemistry
285
24695-706
2010
Show Abstract
Integrin alphavbeta8 is a critical regulator of transforming growth factor beta activation in vasculogenesis during development, immune regulation, and endothelial/epithelial-mesenchymal homeostasis. Recent studies have suggested roles for integrin beta8 in the pathogenesis of chronic obstructive pulmonary disease, brain arteriovenous malformations, and select cancers (Araya, J., Cambier, S., Markovics, J. A., Wolters, P., Jablons, D., Hill, A., Finkbeiner, W., Jones, K., Broaddus, V. C., Sheppard, D., Barzcak, A., Xiao, Y., Erle, D. J., and Nishimura, S. L. (2007) J. Clin. Invest. 117, 3551-3562; Su, H., Kim, H., Pawlikowska, L., Kitamura, H., Shen, F., Cambier, S., Markovics, J., Lawton, M. T., Sidney, S., Bollen, A. W., Kwok, P. Y., Reichardt, L., Young, W. L., Yang, G. Y., and Nishimura, S. L. (2010) Am. J. Pathol. 176, 1018-1027; Culhane, A. C., and Quackenbush, J. (2009) Cancer Res. 69, 7480-7485; Cambier, S., Mu, D. Z., O'Connell, D., Boylen, K., Travis, W., Liu, W. H., Broaddus, V. C., and Nishimura, S. L. (2000) Cancer Res. 60, 7084-7093). Here we report the first identification and characterization of the promoter for ITGB8. We show that a SP binding site and a cyclic AMP response element (CRE) in the ITGB8 core promoter are required for its expression and that Sp1, Sp3, and several AP-1 transcription factors form a complex that binds to these sites in a p38-dependent manner. Furthermore, we demonstrate the requirement for Sp3, ATF-2, and p38 for the transcription and protein expression of integrin beta8. Additionally, reduction of SP3 or inhibition of p38 blocks alphavbeta8-mediated transforming growth factor beta activation. These results place integrin beta8 expression and activity under the control of ubiquitous transcription factors in a stress-activated and pro-inflammatory pathway. Full Text Article | | 20519498
 |
Signature-based small molecule screening identifies cytosine arabinoside as an EWS/FLI modulator in Ewing sarcoma.
Stegmaier, K; Wong, JS; Ross, KN; Chow, KT; Peck, D; Wright, RD; Lessnick, SL; Kung, AL; Golub, TR
PLoS medicine
4
e122
2007
Show Abstract
The presence of tumor-specific mutations in the cancer genome represents a potential opportunity for pharmacologic intervention to therapeutic benefit. Unfortunately, many classes of oncoproteins (e.g., transcription factors) are not amenable to conventional small-molecule screening. Despite the identification of tumor-specific somatic mutations, most cancer therapy still utilizes nonspecific, cytotoxic drugs. One illustrative example is the treatment of Ewing sarcoma. Although the EWS/FLI oncoprotein, present in the vast majority of Ewing tumors, was characterized over ten years ago, it has never been exploited as a target of therapy. Previously, this target has been intractable to modulation with traditional small-molecule library screening approaches. Here we describe a gene expression-based approach to identify compounds that induce a signature of EWS/FLI attenuation. We hypothesize that screening small-molecule libraries highly enriched for FDA-approved drugs will provide a more rapid path to clinical application.A gene expression signature for the EWS/FLI off state was determined with microarray expression profiling of Ewing sarcoma cell lines with EWS/FLI-directed RNA interference. A small-molecule library enriched for FDA-approved drugs was screened with a high-throughput, ligation-mediated amplification assay with a fluorescent, bead-based detection. Screening identified cytosine arabinoside (ARA-C) as a modulator of EWS/FLI. ARA-C reduced EWS/FLI protein abundance and accordingly diminished cell viability and transformation and abrogated tumor growth in a xenograft model. Given the poor outcomes of many patients with Ewing sarcoma and the well-established ARA-C safety profile, clinical trials testing ARA-C are warranted.We demonstrate that a gene expression-based approach to small-molecule library screening can identify, for rapid clinical testing, candidate drugs that modulate previously intractable targets. Furthermore, this is a generic approach that can, in principle, be applied to the identification of modulators of any tumor-associated oncoprotein in the rare pediatric malignancies, but also in the more common adult cancers. | | 17425403
 |
Retinoic acid-induced chromatin remodeling of mouse kappa opioid receptor gene.
Park, SW; Huq, MD; Loh, HH; Wei, LN
The Journal of neuroscience : the official journal of the Society for Neuroscience
25
3350-7
2005
Show Abstract
The mouse kappa opioid receptor (KOR) gene is constitutively expressed in P19 embryonic stem cells but is first suppressed and reactivated during retinoic acid (RA)-induced neuronal differentiation. However, no RA response element (RARE) can be found in this gene regulatory region. The suppression and reactivation of the KOR gene in this neuronal differentiation model suggested chromatin remodeling occurred on this gene promoter triggered by RA induction. This study asks whether RA induces alteration in the nucleosomal structure of this gene promoter that has no apparent RARE and, if so, how RA remodels chromatin of this promoter. The results revealed two loose nucleosomes, N1 at -44 (3' boundary) from the transcription initiation site and N2 spanning the transcription initiation site, that are relevant to active transcription. RA formed a repressive chromatin configuration of this promoter by compacting nucleosome N1, followed by nucleosome N2 condensation. Chromatin immunoprecipitation assay demonstrated RA induced replacement of the c-Myc/Max complex with the Max/Mad1 complex on the E box located within nucleosome N1, coinciding with reduced Sp1 binding to GC boxes located within nucleosome N2 and recruitment of chromatin remodeling factor Brahma-related gene 1 (BRG-1) to this promoter. Consistently, histone deacetylation, Lys9 methylation, and hypophosphorylation of RNA polymerase II C-terminal domain were detected on this promoter after RA treatment. It is concluded that RA induces KOR gene suppression, as early neuronal differentiation marker, by inducing substitution of c-Myc/Max with Max/Mad on the E box and by BRG-1 involved nucleosome recruitment and chromatin condensation, thereby abolishing Sp1 binding. | | 15800190
 |
The cell-specific expression of endothelial nitric-oxide synthase: a role for DNA methylation.
Chan, Y; Fish, JE; D'Abreo, C; Lin, S; Robb, GB; Teichert, AM; Karantzoulis-Fegaras, F; Keightley, A; Steer, BM; Marsden, PA
The Journal of biological chemistry
279
35087-100
2004
Show Abstract
The basis for the endothelial cell-restricted expression of endothelial nitric-oxide synthase (eNOS) is not known. While transgenic promoter/reporter mice demonstrated endothelium cell-specific eNOS expression, we found robust expression of episomal eNOS promoter/reporter constructs in cell types that do not express the native eNOS transcript. To explore the mechanism underlying this differential activity pattern of chromatin-versus episome-based eNOS promoters, we examined the methylation status of 5'-regulatory sequences of the human eNOS gene. DNA methylation differed dramatically between endothelial and nonendothelial cell types, including vascular smooth muscle cells. This same cell type-specific methylation pattern was observed in vivo in endothelial and vascular smooth muscle cells of the mouse aorta at the native murine eNOS promoter. We addressed the functional consequences of methylation on eNOS transcription using transient transfection of in vitro methylated promoter/reporter constructs and found that methylated constructs exhibited a marked decrease in the synergistic action of Sp1, Sp3, and Ets1 on eNOS promoter activity. The addition of methyl-CpG-binding protein 2 further reduced the transcriptional activity of methylated eNOS constructs. Importantly, chromatin immunoprecipitation demonstrated the presence of Sp1, Sp3, and Ets1 at the native eNOS promoter in endothelial cells but not in vascular smooth muscle cells. Finally, robust expression of eNOS mRNA was induced in nonendothelial cell types following inhibition of DNA methyltransferase activity with 5-azacytidine, demonstrating the importance of DNA methylation-mediated repression. This report is the first to show that promoter DNA methylation plays an important role in the cell-specific expression of a constitutively expressed gene in the vascular endothelium. | | 15180995
 |